Introduction
Python may be one of the most popular programming languages today, but it's certainly not the most efficient. Especially in the world of machine learning, users sacrifice efficiency for the ease of use that Python offers.
But that doesn't mean you can't speed things up in other ways. Cython is an easy way to reduce the computational time of Python scripts without sacrificing the functionality that's easily achieved with Python.
This tutorial will introduce you to using Cython to speed up Python scripts. We'll tackle a simple but computationally expensive task: creating a for loop that iterates through a Python list of 1 billion numbers and sums them. Since time is of the essence when running code on resource-constrained devices, we'll explore this issue by considering how to implement Python code in Cython on a Raspberry Pi (RPi). Cython makes a significant difference in computational speed; like a lazy versus a yozo.
Prerequisites for optimizing Python scripts with Cython
- Basic knowledge of Python: Familiarity with Python syntax, functions, data types, and modules.
- Understand basic C/C++ concepts: Familiarity with basic C or C++ concepts such as pointers, data types, and control structures.
- Python development environment: Install Python (preferably Python 3.x) with a package manager like pip.
- Install Cython: Install Cython with the pip install cython command.
- Terminal/Command Line Familiarity: Basic ability to navigate and execute commands in a terminal or command line.
These prerequisites will help you get ready to start optimizing Python code using Cython.
Python and CPython
Many people are unaware of the fact that languages like Python are actually implemented in other languages. For example, the C implementation of Python is known as CPython. Note that this is different from Cython. For more information about the different Python implementations, you can read this article.
The default and most popular implementation of Python is C-Python. Using it has one important advantage. C is a compiled language, and its code is converted to machine code that is executed directly by the central processing unit (CPU). Now, you might ask, if C is a compiled language, does that mean Python is the same?
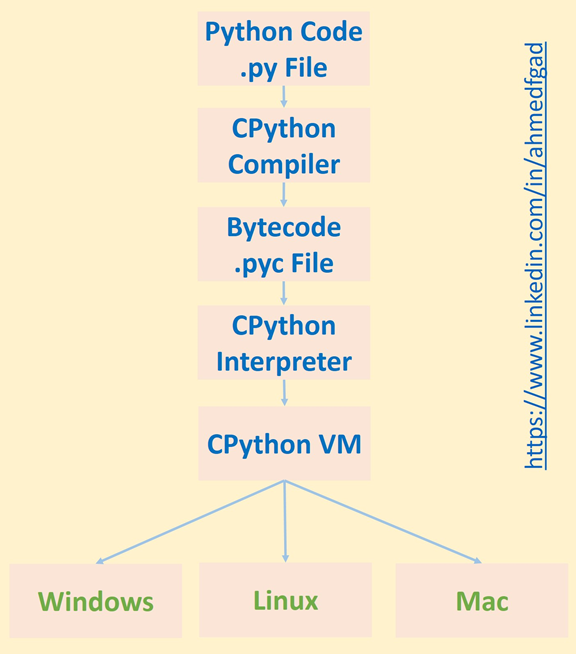
Python implementation in C (Cpython) 100% is not compiled, nor is it interpreted. In fact, both compilation and interpretation occur in the process of executing a Python script. To clarify this, let's look at the steps involved in executing a Python script:
- Compiling source code using CPython to produce bytecode
- Bytecode interpretation by the CPython interpreter
- Running the output of the CPython interpreter in the CPython virtual machine
The compilation process occurs when Cpython compiles the source code (.py file) and produces Cpython bytecode (.pyc file). The Cpython bytecode is then interpreted by the Cpython interpreter and the output is executed in the Cpython virtual machine. As shown in the steps above, the process of executing a Python script involves both compilation and interpretation.
The Cpython compiler produces bytecode only once, but the interpreter is called every time the code is executed. Interpreting bytecode usually takes a long time. If using an interpreter slows down execution, why use it at all? The main reason is that the interpreter makes Python usable on different operating systems. Since the bytecode is executed independently of the machine in the Cpython virtual machine that runs on the CPU, it can be run on different machines without any changes.
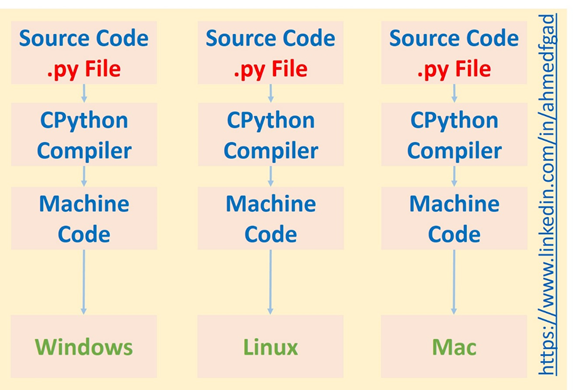
If an interpreter is not used, the Cpython compiler will produce machine code that runs directly on the CPU. Because different platforms have different instructions, the code cannot run on different platforms.
In short, using a compiler makes the process faster, but the interpreter makes the code cross-platform. So, one of the reasons why Python is slower than C is because of the use of the interpreter. Remember that the compiler runs only once, but the interpreter runs every time the code is executed.
Python is much slower than C, but many programmers still prefer it because it is much easier to use. Python hides many details from the programmer, which can prevent frustrating debugging. For example, because Python is a dynamically typed language, there is no need to specify the type of every variable in the code – Python automatically infers it. In contrast, in statically typed languages (such as C, C++, or Java) you must specify the types of variables, as shown below.
int x = 10
string s = "Hello"Compare with the following implementation in Python:
Dynamic typing makes coding easier, but it puts more burden on the machine to find the right data type. This slows down the execution process.
x = 10
s = "Hello"In general, “high-level” languages like Python are much easier for developers to understand. But when the code is executed, it needs to be converted to low-level instructions. This conversion takes more time, which is sacrificed in favor of ease of use.
If time is a factor, you should use low-level commands. So, instead of writing code in Python, which is the front-end, you can write it using CPython, which is Python behind the scenes and implemented in C. However, if you do this, you will feel like you are programming in C, not Python.
CPython is much more complex. In CPython, everything is implemented in C. There is no way to escape the complexity of C when coding. That is why many developers use Cython instead. But how is Cython different from CPython?
How is Cython different?
As defined above, Cython is a language that allows you to have the best of both worlds: speed and ease of use. You can still write regular code in Python, but to speed up execution time, Cython allows you to replace some parts of Python code with C. So, you end up combining both languages in one file. Note that you can imagine that everything in Python is valid in Cython, but with some limitations.
A regular Python file has the extension .py, but a Cython file has the extension .pyx. The same Python code can be written inside .pyx files, but these files also allow you to use Cython code. Note that putting Python code in a .pyx file may make the process faster than running Python code directly, but it won't be as fast as declaring the types of variables. Therefore, the focus of this tutorial is not just on writing Python code inside a .pyx file, but on making modifications that will make it run faster. This adds a bit of complexity to the programming, but it saves a lot of time. If you have some experience with C programming, this will be easier for you.
Citronizing simple Python code
To convert Python code to Cython, you first need to create a file with the extension . .pyx Create instead of extension .py. Inside this file, you can start writing regular Python code (note that there are some limitations on the code that Cython accepts, which are explained in the Cython documentation).
Before proceeding, make sure that Cython is installed. You can do this using the following command.
pip install cython
To generate the .pyd/.so file, we first need to build the Cython file. The .pyd/.so file represents the module that we will later import. A setup.py file is used to build the Cython file. Create this file and put the following code in it. We will use the distutils.core.setup() function to call the Cython.Build.cythonize() function, which will cyanogenize the .pyx file. This function accepts the path to the file you want to cyanogenize. Here, I have assumed that the setup.py file is located in the same location as the test_cython.pyx file.
import distutils.core
import Cython.Build
distutils.core.setup(
ext_modules = Cython.Build.cythonize("test_cython.pyx"))To build the Cython file, enter the following command at the command line. The current directory of the command line is expected to be the same directory as the setup.py file.
python setup.py build_ext --inplace
After this command finishes, two files will be placed next to the .pyx file. The first file will have the extension .c and the other file will have the extension .pyd (or similar, depending on the operating system used). To use the generated file, simply import the test_cython module and the “Hello Cython” message will be displayed directly, as you can see below.
We have now successfully Cytonized the Python code. The next section will look at Cytonizing a .pyx file that has a loop created in it.
Cytonicizing a "for" loop“
Now let's optimize our previous task: a for loop that loops through 1 million numbers and sums them. Let's start by examining the efficiency of just the loop itself. The time module comes in to estimate how long it takes to execute.
import time
t1 = time.time()
for k in range(1000000):
pass
t2 = time.time()
t = t2-t1
print("%.20f" % t)In a .pyx file, the average time for 3 executions is 0.0281 seconds. The code is running on a machine with a Core i7-6500U @ 2.5 GHz processor and 16 GB of DDR3 RAM.
Compare this to the execution time for a typical Python file, which averages 0.0411 seconds. This means that Cython is only 1.46 times faster than Python for iterations, even without having to modify the for loop to run at C speed.
Now let's add the addition. We will use the range() function to do this.
import time
t1 = time.time()
total = 0
for k in range(1000000):
total = total + k
print "Total =", total
t2 = time.time()
t = t2-t1
print("%.100f" % t)Note that both scripts return the same value, which is 499999500000. In Python this takes an average of 0.1183 seconds to run (across three tests). In Cython this is 1.35 times faster, taking an average of 0.0875 seconds.
Now let's look at another example where the loop starts at 0 and goes through 1 billion numbers.
import time
t1 = time.time()
total = 0
for k in range(1000000000):
total = total + k
print "Total =", total
t2 = time.time()
t = t2-t1
print("%.20f" % t)The Cython script finished in about 85 seconds (1.4 minutes), while the Python script finished in about 115 seconds (1.9 minutes). In both cases, it takes a lot of time. What's the point of using Cython if it takes more than a minute to do such a simple task? Note that this is our fault, not Cython's.
As mentioned earlier, writing Python code inside a Cython script (.pyx) is an improvement, but it doesn't make a significant difference in runtime. We need to make some changes to the Python code inside the Cython script. The first thing we need to do is explicitly declare the data type of the variables used.
Assigning C data types to variables
As per the previous code, 5 variables are used: total, k, t1, t2, and t. The data type of all these variables is implicitly inferred by the code, so it takes more time. To save time spent inferring the data type, let's assign the data type from C language.
The type of the variable total is unsigned long long int. It is an integer because the sum of all the numbers is an integer and it is unsigned because the sum will always be positive. But why long long? Because the sum of all the numbers is very large, so long long is used to make the variable as large as possible.
The data type assigned to the variable k is int and the remaining three variables t1, t2, and t are assigned the data type float.
import time
cdef unsigned long long int total
cdef int k
cdef float t1, t2, t
t1 = time.time()
for k in range(1000000000):
total = total + k
print "Total =", total
t2 = time.time()
t = t2-t1
print("%.100f" % t)Notice that the precision defined in the last print statement is set to 100 and all these numbers are zero (see the next image). This is what we can expect from using Cython. While Python takes more than 1.9 minutes, Cython takes no time at all. I wouldn't even say that it is 1000 or 100000 times faster than Python; I tried different precisions for the printed time and still no number appears.
Note that you can also create an integer variable to hold the value passed to the range() function. This will improve performance even further. The new code is below, where the value is stored in the integer variable maxval.
import time
cdef unsigned long long int maxval
cdef unsigned long long int total
cdef int k
cdef float t1, t2, t
maxval=1000000000
t1=time.time()
for k in range(maxval):
total = total + k
print "Total =", total
t2=time.time()
t = t2-t1
print("%.100f" % t)Now that we've seen how to increase the performance of Python scripts using Cython, let's apply this to the Raspberry Pi (RPi).
Raspberry Pi accessibility from a personal computer
If this is your first time using your Raspberry Pi, you will need to connect both your PC and your RPi to a network. You can do this by connecting both devices to a switch that has DHCP (Dynamic Host Configuration Protocol) enabled to automatically assign IP addresses. Once the network is successfully created, you can access your RPi based on the IPv4 address assigned to it. How do you find out what IPv4 address your RPi is assigned to? Don’t worry, you can simply use an IP scanner tool. In this tutorial, I will be using a free app called Advanced IP Scanner.
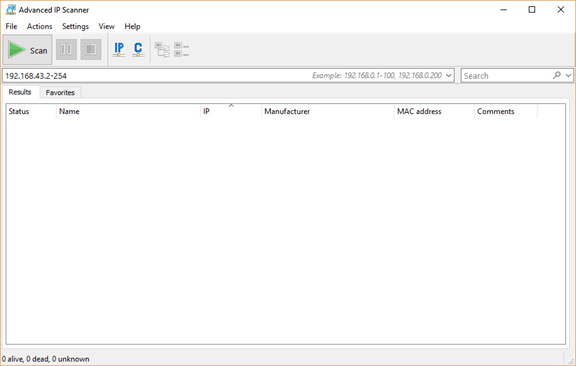
The user interface of this application is as follows. This application accepts a range of IPv4 addresses for searching and returns information about active devices.
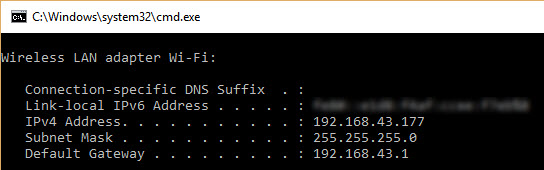
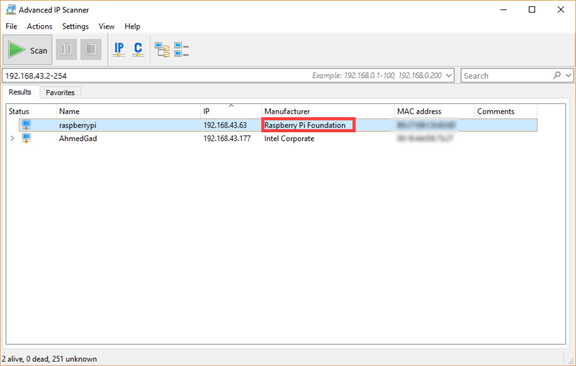
You need to enter the IPv4 address range on your local network. If you don't know this range, just run the ipconfig command in Windows (or ifconfig in Linux) to find your computer's IPv4 address (as shown in the figure below). In my case, the IPv4 address assigned to my computer's Wi-Fi adapter is 192.168.43.177 and the subnet mask is 255.255.255.0. This means that the IPv4 address range on the network is from 192.168.43.1 to 192.168.43.255. As shown in the figure, the IPv4 address 192.168.43.1 is assigned to the Gateway. Note that the last IPv4 address in this range, 192.168.43.255, is reserved for broadcast messages. So, the range you should search starts at 192.168.43.2 and ends at 192.168.43.254.
According to the scan result shown in the next figure, the IPv4 address assigned to the RPi is 192.168.43.63. This IPv4 address can be used to establish a Secure Shell (SSH) session.
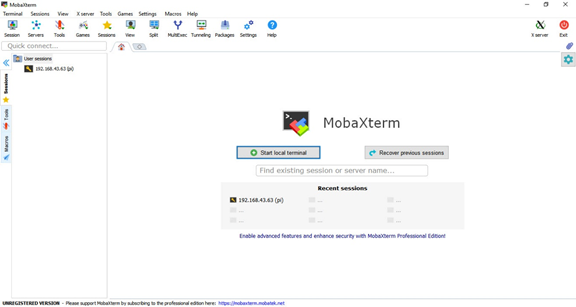
To establish an SSH session, I will use a free software called MobaXterm. The user interface of this program is as follows.
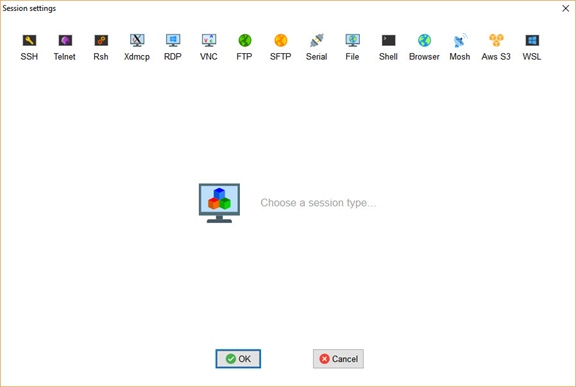
To create an SSH session, simply click on the Session button in the top-left corner. A new window will appear as shown below.
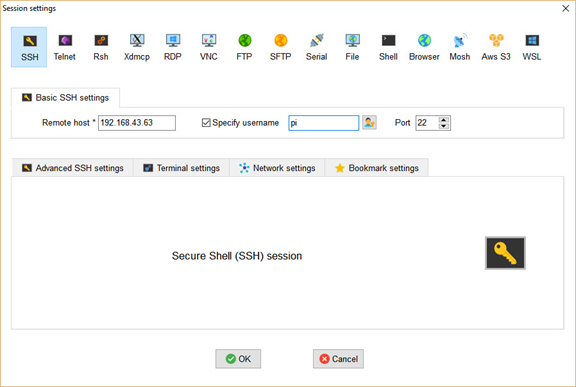
From this window, click the SSH button in the top-left corner to open the window shown below. Simply enter the Raspberry Pi's IPv4 address and username (which is pi by default), then click OK to start the session.
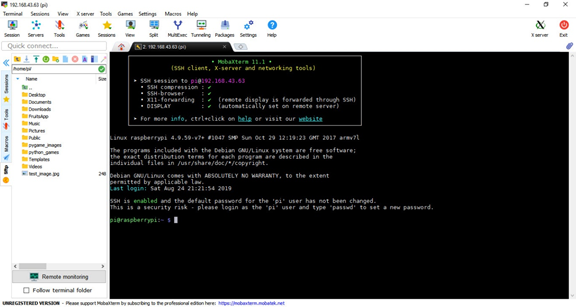
After clicking the OK button, a new window will appear asking you to enter the password. The default password is raspberrypi. After logging in, the following window will appear. The left pane will help you navigate through the directories of your Raspberry Pi easily. There is also a command line to enter commands.
Using Cython with Raspberry Pi
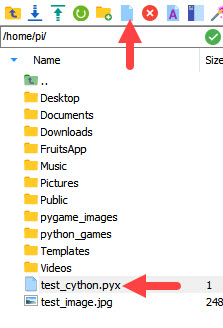
Create a new file and change its extension to .pyx to write the code from the previous example. In the left panel bar there are options for creating new files and directories. You can use the new file icon to make this easier, as shown in the image below. I created a file called test_cython.pyx in the root directory of the Raspberry Pi.
Double click the file to open it, paste the code and save it. Next we need to create the setup.py file which will be exactly as we discussed earlier. After that we need to issue the following command to build the Cython script.
python3 setup.py build_ext --inplace
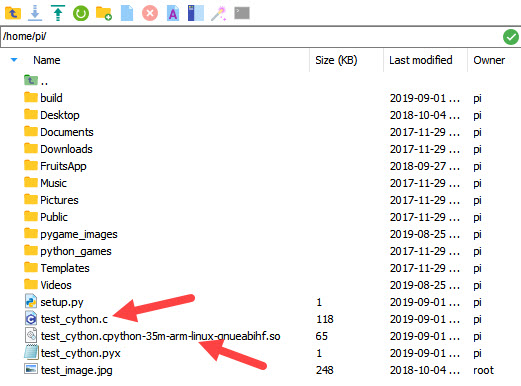
After this command completes successfully, you can find the output files in the left panel, as shown in the following figure. Note that the extension of the module to import is now .so, since we are no longer using Windows.
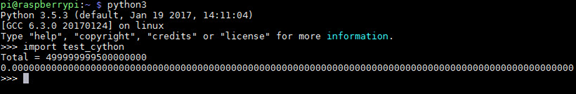
Now let's activate Python and import the module, as shown below. The same results are obtained here as on a PC; the time taken is practically zero.
Result
This tutorial looked at how to use Cython to reduce the computational time of running Python scripts. We will show an example of using a loop. for We looked at adding all the elements of a Python list of 1 billion numbers and compared the execution time with and without declaring the variables. While this process takes nearly two minutes in pure Python, using Cython and declaring static variables, the process runs in virtually no time.